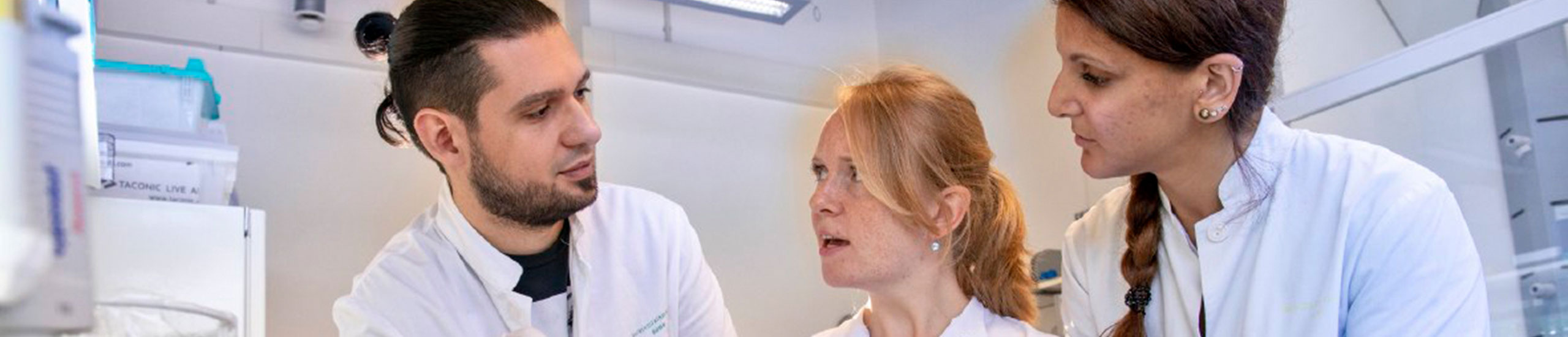
Schmidt Junior Research Group (Immune Monitoring & Genomics)

Dr. rer. nat. Susanne V. Schmidt
Phone: +49 228 287-12102
Fax: +49 228 287-16094
North Zone, Building No. B 24
1st floor, room 1G041
The Immune Monitoring & Genomic (IMG) Group (Schmidt laboratory) seeks to identify biomarkers and develop stratification systems for patients at risk for hyperinflammatory immune responses by using deep immune profiling approaches.
Our laboratory is set up to characterize immune responses on epigenetic, transcriptional and protein levels in the course of diverse inflammatory and rare diseases. We make use of in-house generated immunoassays to track immune responses to danger signals in whole blood, profile cytokine expression patterns using multiplexing bead-based assays and characterize the activation states of monocytes in peripheral blood by flow cytometry. Especially monocytes as major drivers of inflammation are of special interest, as we aim to elaborate molecular mechanisms of adaptive monocyte programming by bioinformatics analysis of transcriptional and epigenetics data.
Clinical Trials
Together with clinical researchers at the medical campus, but also other international collaborators, we aim to identify patients at risk for hyperinflammatory immune diseases, which can lead to severe life-threatening complications during hospitalization or prevent a fast recovery from routine surgeries.
Our scientific mission is to identify people with a hyperinflammatory immune system using an in-vitro whole-blood immune activation assay, which uses various immune triggers to induce inflammatory responses. Utilizing RNA-sequencing techniques and multiplexing assays for classical cytokines present in the blood, we are able to quantify bioinformatically the immune activation status of each patient and are able to predict if the patient is at risk to develop severe complications based on a systemic activation of the immune system, which could be prevented by the administration of anti-inflammatory drugs.
TAM Receptors
The three receptors TYRO3, AXL and MERTK belong to the group of TAM receptors, which are part of the protein tyrosine kinases (PTKs) subfamily. Their activity regulates different critical cellular processes, like apoptosis or growth, but also in the development and progression of various cancer types.
Interestingly, TAM receptors have been shown to play an important role in the homeostasis of the immune system. TAM receptor deficiency or malfunction participates in a broad spectrum of autoimmune diseases, which are caused by the overactivation of antigen-presenting cells and the disability to clear apoptotic debris.
Our current research focuses on the regulation of TAM receptor expression in inflammatory diseases and on mechanisms to increase TAM receptor expression as part of anti-inflammatory therapeutic strategies to support the resolution phase of immune responses and the re-establishment of immune homeostasis.
COVID-19
Our major research focuses currently on the elucidation of immunological pathways, which are modulated by the SARS-CoV-2 virus.
The overreactive immune response in severe COVID-19 patients is characterized by a massive release of cytokines (also called cytokine storm), organ damage and in many cases a compromised recovery from the disease. As monocytes play a central role in hosting immune responses, they are mediators of pro-inflammatory immune responses against SARS-CoV-2 but are also involved in an unrestrained, ongoing immune activation in severe COVID-19 disease courses. Using RNA-sequencing technologies and ChIP-sequencing for classical histone modifications, we track changes in the transcriptome and epigenome of monocytes over the course of SARS-CoV-2 infection and correlate it to the activation stage and changes in biological functions of monocytes.
By the integration of genomics data, clinical patient and laboratory data via bioinformatics analysis, we aim to understand molecular mechanisms and immunological circuits which lead to an ongoing, unrestrained immune response in severe COVID-19 diseases courses, while other patients show only mild or even no symptoms of infection. Our research will hopefully contribute to an improved understanding of COVID-19 and identify patients at risk for severe COVID-19 disease courses and potential long-COVID patients.
Immunogenomics
Our current research is focused on the discovery of dysregulated or malfunctioning programs of myeloid cells in inflammatory diseases and biomarker discovery for therapeutic approaches. Our special interest and core expertise focus on transcriptional and epigenetic programming of myeloid cells like monocytes, macrophages and dendritic cells.
Multi-omics approaches enable us to draw a clear picture of the transcriptional process in inflammatory myeloid cells, the underlying epigenetic mechanisms and the resulting biological functions of myeloid cells.
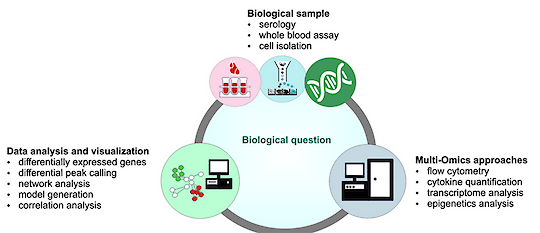
In combination with additional data on patient history, cytokine expression (bead-based multiplexing techniques) and surface marker expression (flow cytometry), we are able to identify central factors, which participate in disease pathogenicity and are potential targets for therapeutic interventions. Following the concept of Systems Biology, we use Next-Generation-Sequencing and Multiplexing-Techniques (mRNA 3'-prime sequencing, ATAC-sequencing and ChIP-sequencing) to generate disease-specific datasets which are analysed in unbiased bioinformatics approaches. Our laboratory owns PC systems with high computing power to analyze each day new incoming data on epigenetic and transcriptional information.
Besides our own research, we support the research of local and international collaborators through bioinformatics analysis, data mining and hypothesis generation.
Group members
Carolyn Krause, Doctoral candidate
Phone: +49 228 287-12118
Marie Rüthing, Doctoral candidate
Phone: +49 228 287-12118
Dr. Jasper Spitzer, PostDoc
Tel.: +49 228 287-12117
Ulrike Strube, Lab technician
Tel.: +49 228 287-12116

(from left to right: Eva Bergmann, Susanne V. Schmidt, Ph.D., Jasper Spitzer, Carolyn Krause, Ulrike Strube)