Research
Our research focuses mainly on two main topics dealing with the analysis of molecular mechanismisms leading to autoimmune diseases (ADs) and the molecular mechanisms during viral recognition. In both processes cytoplasmic RNA sensors, namely RIG-I like receptors (RLRs), play a crucial role. Their role is more obvious in virus recognition since RLRs distinguish between viral RNAs and self-RNAs and trigger type I interferons (type I IFNs) as a defense mechanism. But it has also been found, that mutations in RLR lead to the development of several autoimmune diseases.

Prof. Dr. Hiroki Kato
Autoimmune disease
 Currently more than 100 different autoimmune diseases (ADs) are known, affecting 10% of the european and 8% of the american human population. The most frequent ADs are rheumatoidic arthritis (RA), systemic lupus erythematosus (SLE), multiple sclerosis (MS), type 1 diabetes (T1D), psoriasis and celiac disease. The reasons for developing a AD are multifactorial and include genetic predisposition, environmental factors and the individual way of living.
Currently more than 100 different autoimmune diseases (ADs) are known, affecting 10% of the european and 8% of the american human population. The most frequent ADs are rheumatoidic arthritis (RA), systemic lupus erythematosus (SLE), multiple sclerosis (MS), type 1 diabetes (T1D), psoriasis and celiac disease. The reasons for developing a AD are multifactorial and include genetic predisposition, environmental factors and the individual way of living.
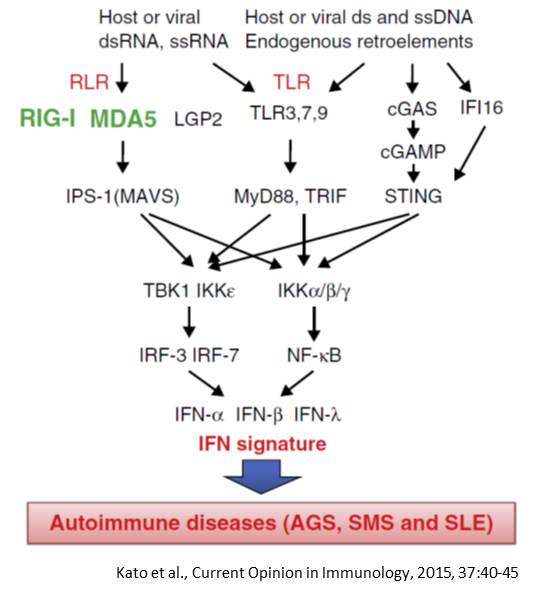
Amongst the genetically caused ADs is a group of disorders having an abnormal upregulation of type I IFN in common and are therefore called interferonopathies. They comprise diseases like SLE, Singleton-Merten Syndrom (SMS), Aicardi-Goutieres Syndrom (AGS) und T1D. Since many ADs and interferonopathies are based on similar or the same genetical cause, it is important to investigate overlapping molecular mechanisms.
Two key molecules in interferonopathies are retinoic acid Inducible Gene I (RIG-I) and Melanoma Differentiation Associated Gene 5 (MDA5). Mutations in both corresponding genes (DDX58 respecively IFIH1) could be connected to the development of ADs and interferonopathies like SLE, SMS, AGS, T1D, MS and psoriasis.
RIG-I and MDA5 are intracellular receptors of viral nucleic acids. When a virus infects a cell, it injects its genetical material and abuses the cellintrinsic machenery for protein production and viral amplification. The viral genetical material is recognized by RIG-I and MDA5 as foreign material and an immune reaction is initiated. Via several signaling molecules within the cells, the production of pro-inflammatory interferons is activated. These are secreted and taken up by surrounding cells via specific receptors. Up on this uptake a chain reaction of signaling molecules is activated which trigger more than 3000 genes of the immune system to fight the infection. Misregulations within this process can lead to an over reacting immune response yielding in ADs and interferonopathies like SLE, SMS, AGS and T1D.
Viral recognition
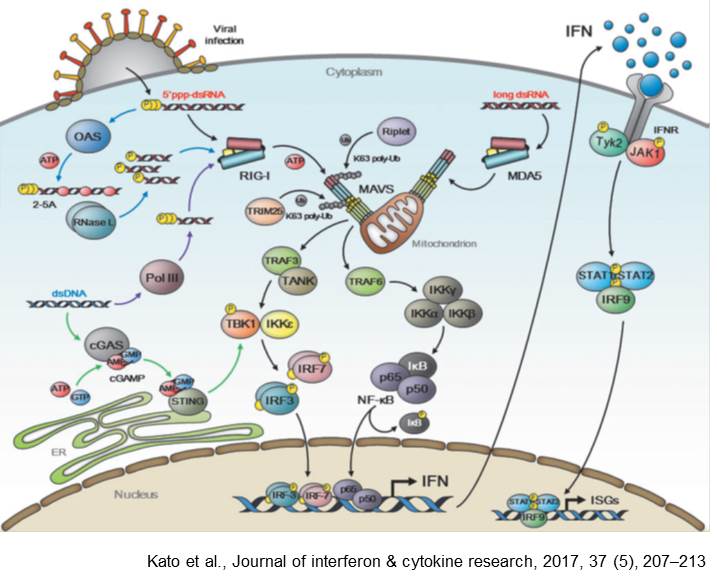 Virus destruction by a cell needs specific receptors, which recognize viral-specific components and initiate an appropriate immune response. Amongst these components are for example double-stranded RNA molecules (dsRNA) which do not appear in normal and healthy cells and are therefore categorized as foreign material.
Virus destruction by a cell needs specific receptors, which recognize viral-specific components and initiate an appropriate immune response. Amongst these components are for example double-stranded RNA molecules (dsRNA) which do not appear in normal and healthy cells and are therefore categorized as foreign material.
The differentiation is done by specific receptors like RIG-I and MDA5. Although both molecules spot dsRNA, it was shown that they still comprise virus-specificity. Whilst RIG-I perceive paramyxo viruses, influenza viruses and viruses of japanese encephalitis, MDA5 recognizes picorna viruses. Furthermore, it was proven that RIG-I and MDA5 are crucial for effective destruction of these viruses during infection.
Upon viral recognition by RIG-I or MDA5 a signaling cascade is initiated resulting in the production of pro-inflammatory cytokines and chemokines. Within these signaling cascades IPS-1 (also known as MAVS) is a central molecule, combining signals from MDA5 and RIG-I into one common signaling chain.
In the end IRF3 and IRF7 are activated and cytokines like tumor necrosis factor alpha (TNF-a), IFN-alpha and IFN-beta are produced. The activation of these signaling cascades has to be tightly regulated to initiate an effective immune response without yielding in a overproduction of cytokines which would result in a deadly so called cytokine storm.
Early virus-host interactions are an interplay of viral exploitation of cellular proteins, functions, and innate defence mechanisms. Our research aims at answering the following questions: Which mechanisms and specific patterns lead to the recognition of a viral infection? What does a virus do to evade this recognition? Which cellular structures resemble viruses? In which cells are viral infections actually recognized – the non-infected, or the infected ones? We are taking a closer look on the constant struggle between viruses and the immune system, with special emphasis on extracellular vesicles (EVs). In recent years, this new and highly interdisciplinary field has led to many surprising discoveries and shows a high potential for further scientific developments.
Please find more information about our research on the website of "Cellular Virology" team.


















